Tutorial
This tutorial walks you through the full TridentSynth workflow using a heptane (C7H16) example with PKS + biological pathways. Whether you work with polyketide synthases, organic chemistry, or computational biology, no prior PKS knowledge is needed to follow along.
Start by telling TridentSynth which molecule you want to synthesize. There are two ways to do this:
Option A: Type a SMILES string
Enter a i
SMILES string directly into the text field.
For example, heptane is represented as: CCCCCCC
Option B: Draw the structure
Click "Draw structure" to open the Ketcher molecule editor, where you can draw your target molecule graphically. When you save, the SMILES string is automatically extracted and filled into the text field.
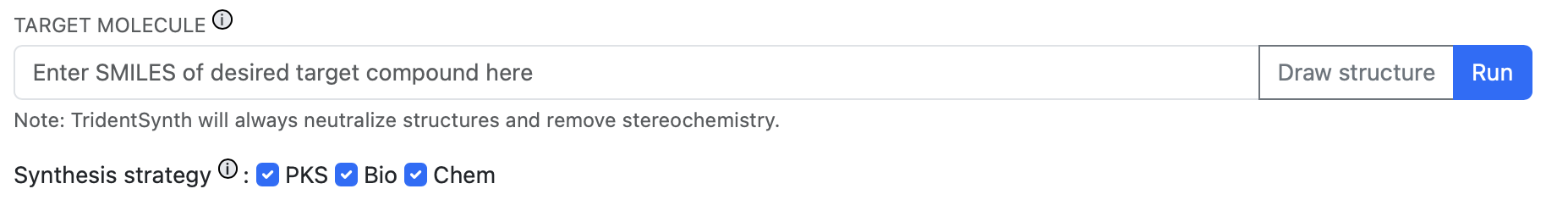
The query form: enter a SMILES string or draw a structure, then select your synthesis strategies.
TridentSynth offers three synthesis routes that can be used individually or in combination. Select one or more checkboxes on the query page:
| Strategy | What it does | Best for |
|---|---|---|
| PKS | Uses type I modular polyketide synthases to build a carbon backbone forward toward the target by stitching together acyl-CoA building blocks. Designs are generated by RetroTide. | Molecules with repeating C2 units, long carbon chains, polyketide-like scaffolds |
| Bio | Uses monofunctional enzyme reaction rules. In PKS+Bio mode, works forward from the PKS product toward the target. In Bio-only mode, works backward from the target to known precursors. Powered by DORAnet. | Functional group modifications, hydroxylations, oxidations, enzymatic tailoring |
| Chem | Uses synthetic organic chemistry reaction rules. Same directional logic as Bio (forward after PKS, backward otherwise). | Transformations not easily achieved by enzymes, broader reaction diversity |
Common combinations
| PKS + Bio | Build the backbone with PKSs, then tailor with enzymes. Used in this tutorial. |
| PKS + Bio + Chem | Maximum search space — combines all three routes |
| PKS only | Quick check if the target is directly reachable by PKS assembly alone |
| Bio only | Find enzymatic retrosynthesis pathways from target back to common metabolites |
| Chem only | Find synthetic chemistry retrosynthesis pathways from target back to common metabolites |
Click "Advanced (Optional)" on the query page to expand additional settings. The defaults work well for most cases, but here is what each parameter controls:
Max biological / chemical synthesis steps
Controls how many successive reaction steps DORAnet will explore when tailoring the PKS product (or when working backward from the target in non-PKS mode). Range: 1–3 steps. More steps broaden the search but increase runtime.
PKS release mechanism
Determines how the polyketide chain is released from the PKS enzyme at the end of assembly:
- Thiolysis — Cleaves the thioester bond to release a free carboxylic acid. This is the most common mechanism and is used in this tutorial.
- Cyclization — Forms an intramolecular ring (lactone) by attacking an internal hydroxyl or amine. Choose this for macrolide-like targets.
PKS starters
The acyl-CoA substrate loaded onto the first (loading) module of the PKS. Available options include malonyl-CoA, methylmalonyl-CoA, ethylmalonyl-CoA, and others. Selecting more starters increases the design space that RetroTide explores.
PKS extenders
The acyl-CoA substrates used by extension modules to elongate the polyketide chain. Each extension module adds a two-carbon unit to the growing chain, with the specific extender determining the side-chain substituent.
Max atoms filter (DORAnet)
Optional upper limits on the number of C, N, and O atoms in intermediate metabolites during Bio/Chem network expansion. This can speed up computation by pruning intermediates that are too large. Leave blank to allow intermediates of any size.
Click "Run" to submit your synthesis job. Here is what happens behind the scenes:
- Validation — Your SMILES is checked for correctness, sanitized with RDKit, and screened against a controlled substances list.
- Job queued — The synthesis task is sent to a background worker (Celery). You receive a unique job ID and a bookmarkable link to your results.
- PKS design (RetroTide) — If PKS is selected, RetroTide combinatorially explores module designs to build a carbon backbone that is structurally similar to your target.
- Post-PKS tailoring (DORAnet) — If Bio and/or Chem is selected, DORAnet searches for reaction pathways to convert the PKS product into the final target.
- Feasibility scoring — Each predicted reaction is scored by an XGBoost feasibility classifier trained on known enzymatic reactions.
- Enzyme retrieval — For enzymatic reactions, TridentSynth searches a database of ~55,000 known reactions to find closely-matched natural enzymes.
Runtime depends on the complexity of your target and the strategies selected. PKS-only jobs typically complete in under a minute; adding Bio/Chem steps with higher step counts may take several minutes.
The results page presents your synthesis designs. Here is a guide to each section:
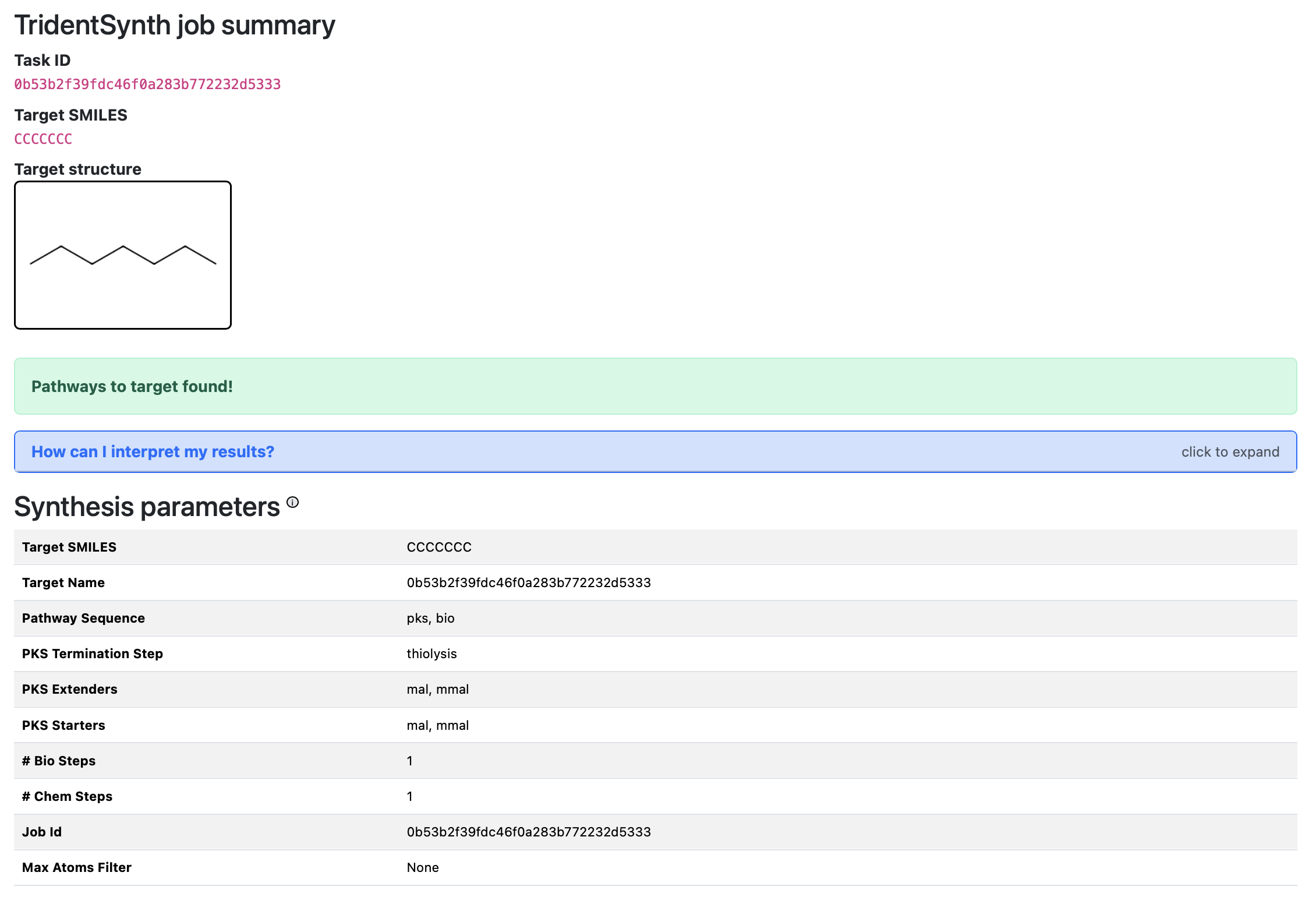
The job summary panel: target structure, status alert, and synthesis parameters used for the run.
Status alerts
A colored banner at the top tells you the outcome:
- Green — Full pathways to your exact target were found
- Yellow — The exact target was not reached, but pathways to the most structurally similar product are shown
- Blue — The target can be synthesized by PKS assembly alone (no post-PKS tailoring needed)
PKS module designs
If PKS was selected, each design shows the engineered PKS assembly as a series of modules. Each module contains catalytic domains that determine how the polyketide chain is modified at that step:
| Domain | Role |
|---|---|
| KS / KSq | Ketosynthase — catalyzes the condensation reaction that extends the chain by two carbons |
| AT | Acyltransferase — selects and loads the extender substrate (e.g., malonyl-CoA) |
| KR | Ketoreductase — reduces the ketone to a hydroxyl group (with stereospecificity: A, B, or C type) |
| DH | Dehydratase — removes water to form a double bond (requires an active KR) |
| ER | Enoylreductase — reduces the double bond to a saturated bond (requires an active DH) |
| ACP | Acyl carrier protein — tethers the growing chain (present in every module) |
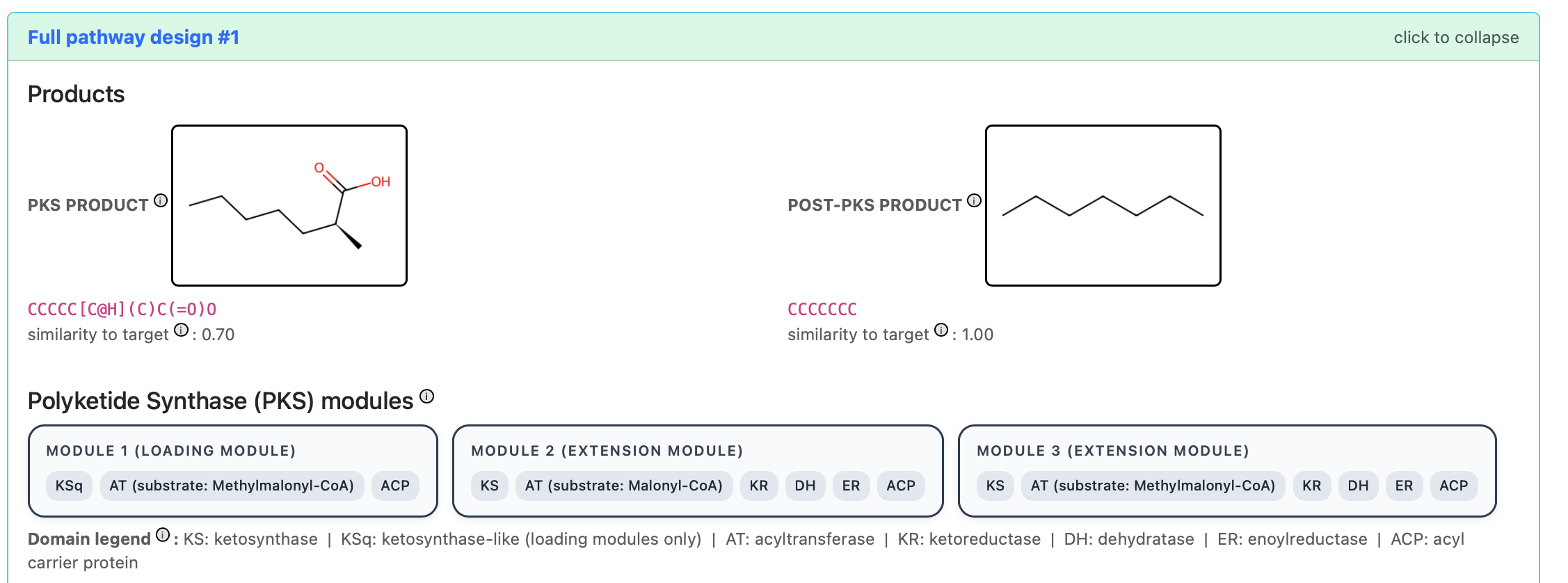
A pathway design: the PKS product (left), post-PKS product (right), and the PKS module architecture with domain annotations.
Similarity scores
Each design reports how structurally similar its product is to your target, measured using the i maximum common substructure (MCS) metric on a 0–1 scale. A score of 1.0 means the product is an exact structural match to the target. Designs are sorted by similarity, with the best matches shown first.
Post-PKS pathways
If Bio and/or Chem steps were used, the results show the predicted reaction pathways connecting the PKS product to the final target (or to the closest reachable product). Each pathway lists:
- Reaction SMILES — The chemical transformation at each step, shown as both a rendered image and a text string
- Reaction rules — The named rule that predicted each reaction (e.g.,
rule0001for enzymatic rules). Enzymatic rules link to their associated UniProt enzyme entries. - Feasibility scores — Each reaction is scored 0–1 by an XGBoost classifier. The net feasibility is the product of all individual scores — higher is better.
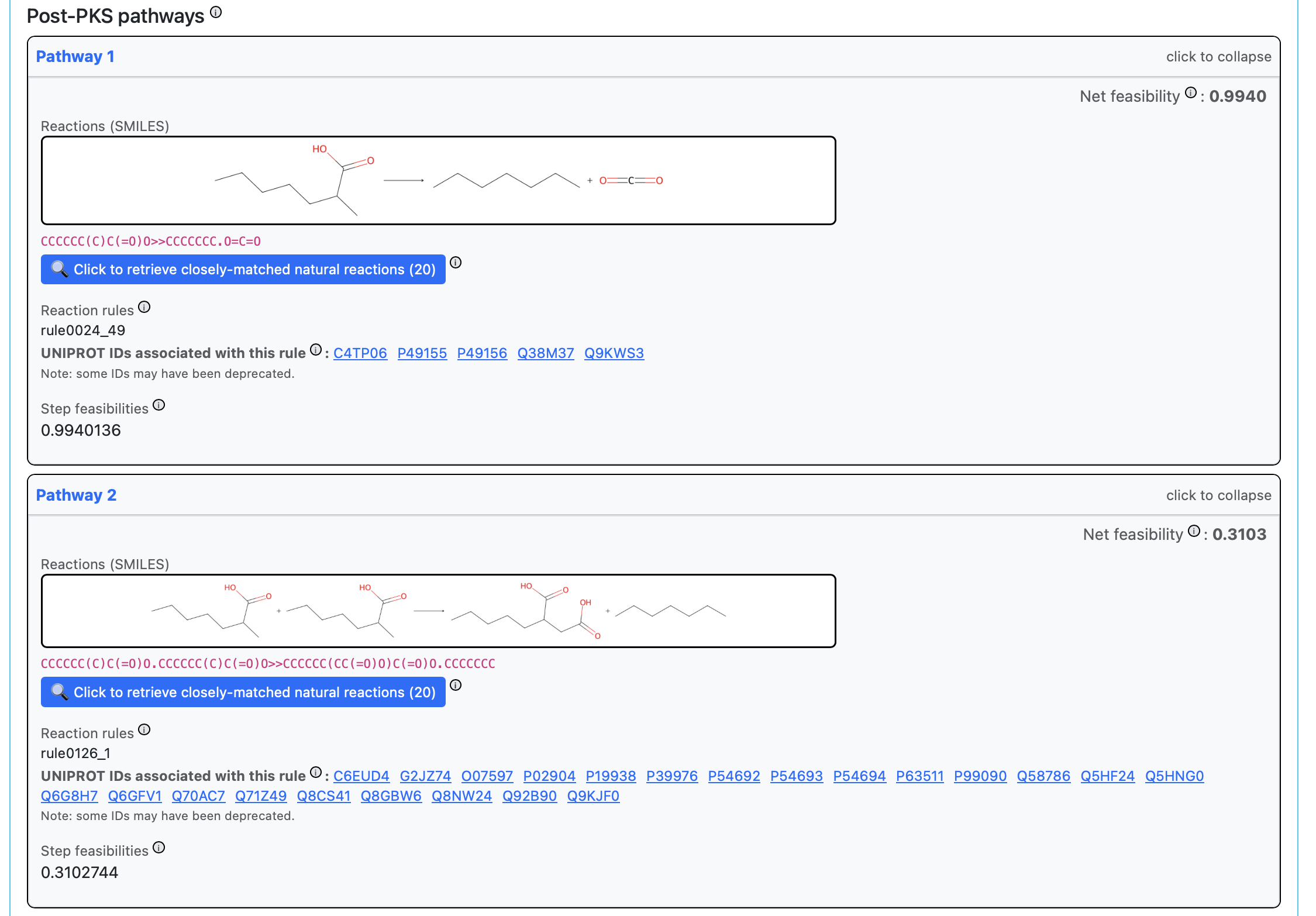
Post-PKS pathways: reaction images, reaction rules with UniProt links, feasibility scores, and enzyme retrieval buttons.
Enzyme retrieval
For each predicted reaction, you can click the search button to retrieve closely-matched natural reactions from a database of ~55,000 known enzymatic reactions. Matches are ranked by i RCMFP similarity and include:
- A rendered image of the matched natural reaction
- The EC number with links to BRENDA, KEGG, and MetaCyc
- Protein reference links to UniProt or GenBank
These matched enzymes can serve as starting points for experimental validation of the predicted pathway.
Ready to try the workflow? Use the buttons below to either load the heptane example into the query form (so you can explore the settings and submit it yourself) or jump straight to the pre-computed results.
Further reading
- About / Help — Detailed background on TridentSynth's algorithms and design
- Publications — Papers to cite when using TridentSynth
- Common precursors — The ~300 metabolites used as targets in non-PKS retrosynthesis